David E. Graff, Eugene I. Shakhnovich, and Connor W. Coley (2020)
Highlighted by
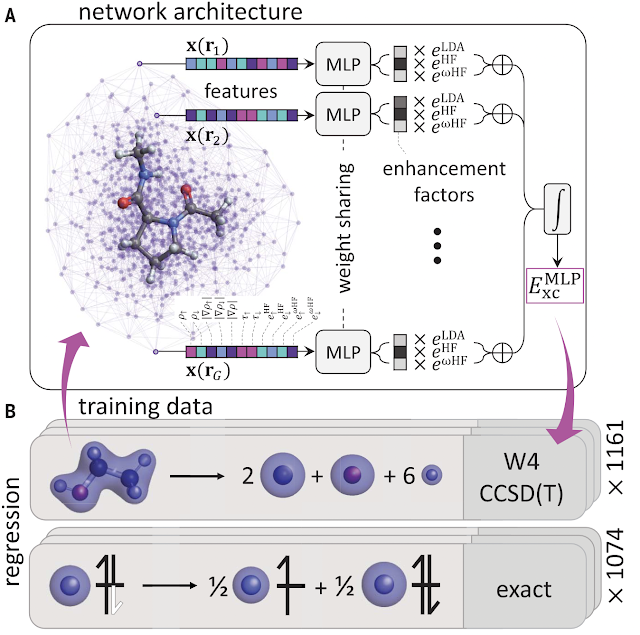
Jan JensenFigure 1 and part of Figure 2 from the paper. (c) The authors 2021.
This paper shows how to find the highest scoring molecules in a very large library of molecules by scoring only a very small percentage of the library. The focus of the paper is docking scores but it can in principle to be used for any molecular property.
The general approach is simple:
1. Start by picking a random sample of the library (say 100 molecules out of a library of 10.000 molecules) and evaluate their scores.
2. Use these 100 points to train a machine-learning (ML) model to predict the scores.
3. Screen all 10,000 molecules using the ML model. The assumption is that training/using the ML model is much cheaper than evaluating the score.
4. Select the 100 best molecules according to the ML model, compute the scores, and use them to retrain the ML model.
5. Repeat steps 3 and 4.
The best molecules could be the best-scoring molecules (this a known as "greedy" optimisation). However, if the uncertainty of the ML prediction for each molecule can be quantified, there are several other options for what best is (use of these approaches are referred to as Bayesian optimisation). The study investigates four selection functions involving standard deviations but finds the greedy approach works best.
The approach is tested on three different datasets with known docking scores of varying sizes (10K, 50K, 2M, and 99M). The study tests three different machine learning models: RF and NN using fingerprints as well as a graph convolutional model (which works best) and various choices batch sizes.
In the case of the 99M dataset more than half of the top-50,000 scoring molecules can be found by docking only 600K molecules using this approach.
However, let's turn that last sentence around: if you're developing an ML model to find high-scoring molecules your training set size needs to be 600K. Furthermore, the study shows that if you just pick 500K random molecules for your training set, your ML model won't identify any of the top-50,0000 molecules. You have to build this very large training set in this iterative fashion to get an ML model that can reliably identify the top-scoring molecules.