Stefan Grimme, Christoph Bannwarth, and Philip Shushkov (2017)
Highlighted by Jan Jensen
Inspired by the success of the sTDA-xTB method for predicting electronic spectra, Grimme and co-workers present the tight binding DFT method GFN-xTB for predicting Geometries, vibrational Frequencies and Noncovalent interactions ("x" stands for extensions in the AO basis set and the form of the Hamiltonian).

This work is licensed under a Creative Commons Attribution 4.0 International License.
Highlighted by Jan Jensen
Copyright American Chemical Society (2017)
Inspired by the success of the sTDA-xTB method for predicting electronic spectra, Grimme and co-workers present the tight binding DFT method GFN-xTB for predicting Geometries, vibrational Frequencies and Noncovalent interactions ("x" stands for extensions in the AO basis set and the form of the Hamiltonian).
The method is implemented in a new program called xtb which includes shared memory parallelization, angeometry optimizer, MD, conformational searches, reaction path optimisation modules, and a GB solvation model supplemented with analytical nuclear gradients. The software is free to academics and can be obtained by writing to the authors.
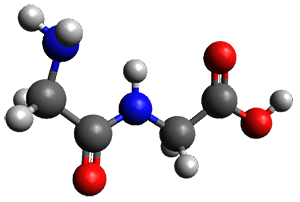
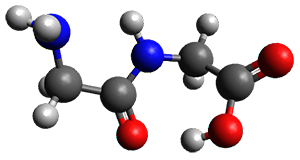
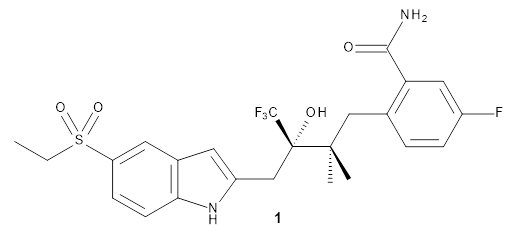
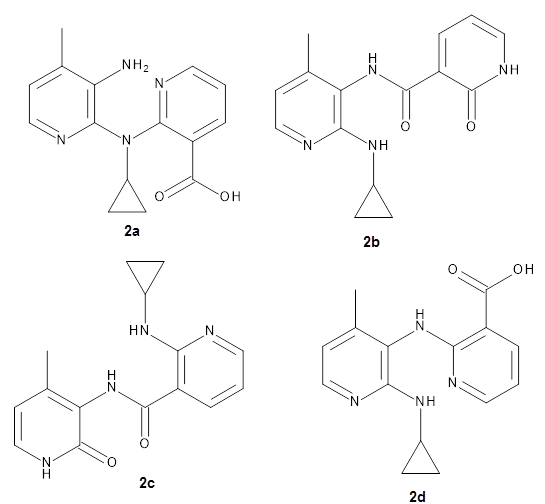
Importantly, the method is parameterized for all spd elements, so finally there is an alternative to PM6 for heavier elements. A Fermi smearing technique helps convergence for molecules with small HOMO-LUMO gaps and GFN-xTB gives structures organometallic compounds that are in good agreement with DFT.
As implied by the name, GFN-xTB is not parameterized against reaction energies but the methods gives reasonable results for properties like heats of formation. However, as the authors write
A key premise of the present method is its special purpose character. In our view, low-cost semiempirical QM methods cannot describe simultaneously very different chemical proper- ties, such as structures and chemical reaction energies, and the GFN-xTB method (as the name conveys) focuses on structural properties. The efficiently computed structures and vibrational frequencies or the conformers obtained from global search procedures can (and should) be used subsequently for more accurate DFT or WFT refinements. We hope that the method can serve as a general tool in quantum chemistry and in particular recommend GFN-xTB optimized structures (and thermostatistical corrections) in a multilevel scheme together with PBEh-3c single-point energies. Large-scale molecular dynamics, screening of huge molecular spaces (libraries), parametrization of force-fields, or providing input for novel machine learning techniques are obvious other fields of application.I thank Anders Christensen for bringing this paper to my attention

This work is licensed under a Creative Commons Attribution 4.0 International License.