Contributed by Steven Bachrach.
Reposted from Computational Organic Chemistry with permission
With the proliferation of density functionals, selecting the functional to use in your particular application requires some care. That is why there have been quite a number of benchmark studies (see these posts for some examples). Yu and Karton have now added to our benchmark catalog with a study of π-conjugation.1
They looked at a set of 60 reactions which involve a reactant with π-conjugation and a product which lacks conjugation. A few examples, showing examples involving linear and cyclic systems, are shown in Scheme 1.
Scheme 1.
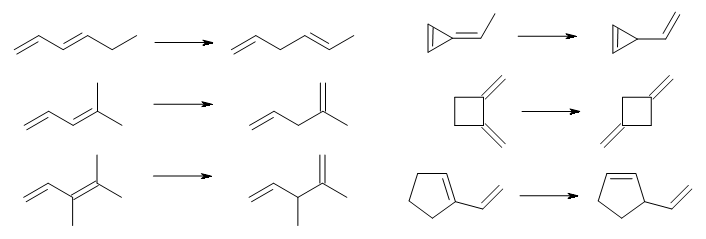
The reaction energies were evaluated at W2-F12, which should have an error of a fraction of a kcal mol-1. Three of the reactions can be compared with experimental values, and difference in the experimental and computed values are well within the error bars of the experiment. It is too bad that the authors did not also examine 1,3-cyclohexadiene → 1,4-cyclohexadiene, a reaction that is both of broader interest than many of the ones included in the test set and can also be compared with experiment.
These 60 reactions were then evaluated with a slew of functionals from every rung of Jacob’s ladder. The highlights of this benchmark study are that most GGA and meta-GGA and hybrid functionals (like B3LYP) have errors that exceed chemical accuracy (about 1 kcal mol-1). However, the range-separated functionals give very good energies, including ωB97X-D. The best results are provided with double hybrid functionals. Lastly, the D3 dispersion correction does generally improve energies by 10-20%. On the wavefunction side, SCS-MPs gives excellent results, and may be one of the best choices when considering computational resources.
References
(1) Yu, L.-J.; Karton, A. "Assessment of theoretical procedures for a diverse set of isomerization reactions involving double-bond migration in conjugated dienes," Chem. Phys. 2014, 441, 166-177, DOI:10.1016/j.chemphys.2014.07.015.

This work is licensed under a Creative Commons Attribution-NoDerivs 3.0 Unported License.
No comments:
Post a Comment